Example animations of HD-EEG topographic maps
example-animations.RmdThis article introduces how to create animated HD-EEG topographic
maps using the diegr package. We will demonstrate how to
visualize changes in EEG amplitude over time using built-in animation
functions.
Topographic map animations in 2D
To display changes in EEG amplitude over time as an animated 2D
topographic map, the diegr package provides two main
functions:
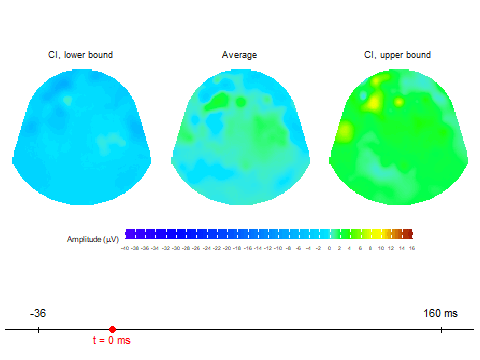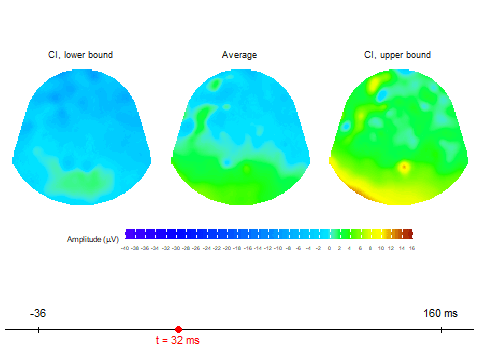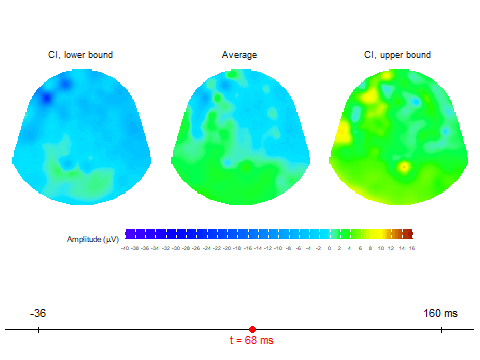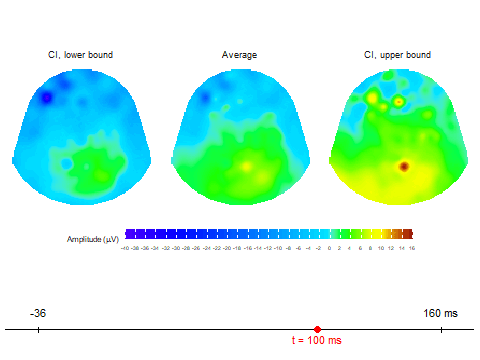
animate_topoCreates basic animated topographic map of a selected variable. The input can be raw EEG signal, averaged signal, or any other amplitude column in the input data frame.animate_topo_meanDesigned specifically for data frames produced by thecompute_meanfunction. In addition to the average signal, this function also visualizes the lower and upper confidence interval bounds.
Example: Animating EEG amplitude after stimulus
In this example, we want to examine the development of EEG amplitude for one subject within a 100 ms time window after stimulus onset.
Data used in this example are built-in dataset
epochdata, we select Subject 1 and the time interval 10
(stimulus onset) to 35 (100 ms after stimulus).
Step 1 – Filtering and cleaning the data
In this example, we will start by inspecting the raw epoched data for potential outliers; we want to identify and remove extreme values before proceeding with further analyses.
For this purpose, we examine response times and amplitude values (for an individual sensor within a selected time range) using interactive boxplots.
# load datasets
data("epochdata")
data("rtdata")
# boxplots of response times
boxplot_rt(rtdata)
# filter subject 1
subject01 <- pick_data(epochdata, subject_rg = 1)
# boxplots of amplitude for sensor E152
pick_data(subject01, sensor_rg = "E152") |>
boxplot_epoch(amplitude = "signal", time_lim = 10:20)From the boxplots, we identify: - epoch 1 as an outlier in response times - epoch 14 as an outlier in amplitude. These two epoch should be excluded before further computation.
# extract outlier epochs
subject01_clean <- pick_data(subject01, epoch_rg = 2:13)At this point, subject01_clean contains data for Subject
1 with the identified outlier epochs removed and is ready for subsequent
analysis steps.
Step 2 - Baseline correction and averaging
Once outliers have been excluded, we need to normalize the signal.
This is done using the baseline_correction function, where
we specify the time range to be used as a reference.
After baseline correction we compute the mean amplitudes for the time window of interest (10 to 35 time points).
# baseline correction
subject01_base <- baseline_correction(subject01_clean, baseline_range = 1:10)
# compute average for selected times
subject01_mean <- subject01_base |>
pick_data(time_rg = 10:35) |>
compute_mean(amplitude = "signal_base", domain = "space",
level = "epoch", type = "jack")At this point, subject01_mean contains averaged data for
Subject 1 within the selected time window and is ready for
animations.
Step 3 – Animation
To provide a clear overview, the animations include a timeline with a
marker indicating the current time. Using the t0 and
t_lim arguments, we can define both the “zero point” of the
animation (e.g., stimulus onset) and the total range of the axis.
Note: Both arguments must be entered as time point indices, not in
miliseconds (e.g., use t0 = 10 instead of
t0 = 0ms).
In addition to rendering within the R viewer, functions also enable
saving animations in gif format to a chosen location.
In this example, we set: t0 = 10 – stimulus onset is at
time point 10t_lim = c(1,50) – the whole time interval presented in the
epochdataFS = 250 – sampling frequency is 250 Hz (i.e. there is
interval 4 ms between two time points)
template = "HCGSN256" – we want plot according to the
HCGSN256 template, dense point mesh is constructed automatically inside
the function
Animation via animate_topo function
animate_topo(subject01_mean, amplitude = "average",
t_lim = c(1,50), t0 = 10, FS = 250,
template = "HCGSN256")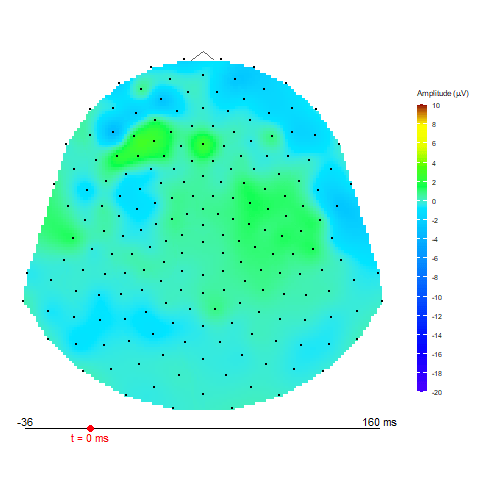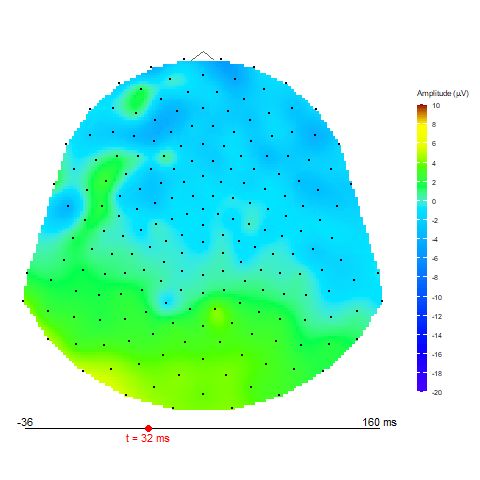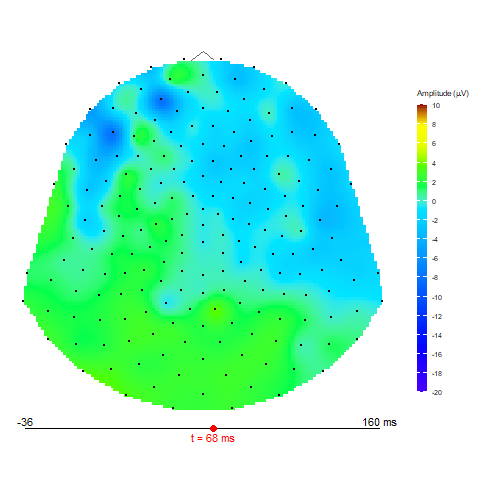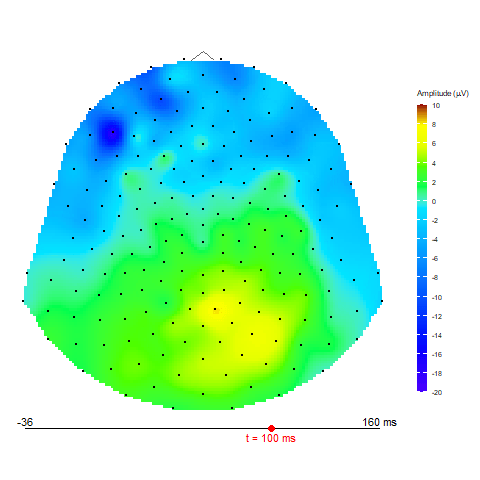
Animation via animate_topo_mean
function
animate_topo_mean(subject01_mean,
t_lim = c(1,50), t0 = 10, FS = 250,
template = "HCGSN256")